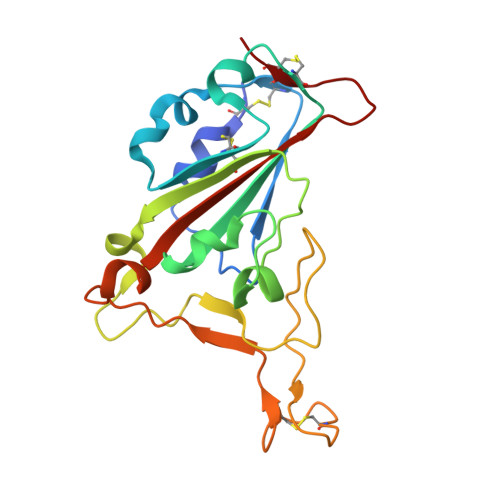
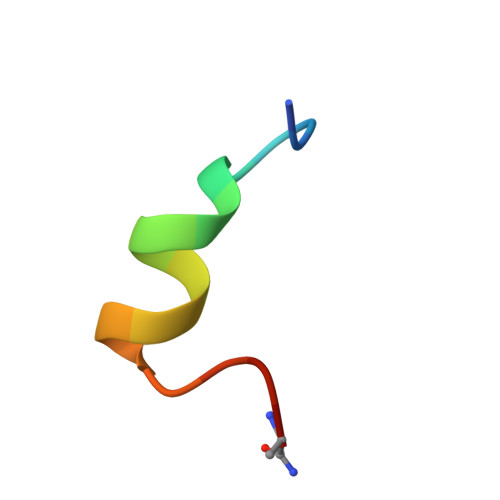
Multivalent bicyclic peptides are an effective antiviral modality that can potently inhibit SARS-CoV-2.
Gaynor, K.U., Vaysburd, M., Harman, M.A.J., Albecka, A., Jeffrey, P., Beswick, P., Papa, G., Chen, L., Mallery, D., McGuinness, B., Van Rietschoten, K., Stanway, S., Brear, P., Lulla, A., Ciazynska, K., Chang, V.T., Sharp, J., Neary, M., Box, H., Herriott, J., Kijak, E., Tatham, L., Bentley, E.G., Sharma, P., Kirby, A., Han, X., Stewart, J.P., Owen, A., Briggs, J.A.G., Hyvonen, M., Skynner, M.J., James, L.C.(2023) Nat Commun 14: 3583-3583
- PubMed: 37328472 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-39158-1
- Primary Citation Related Structures:
7Z8O, 8AAA - PubMed Abstract:
COVID-19 has stimulated the rapid development of new antibody and small molecule therapeutics to inhibit SARS-CoV-2 infection. Here we describe a third antiviral modality that combines the drug-like advantages of both. Bicycles are entropically constrained peptides stabilized by a central chemical scaffold into a bi-cyclic structure. Rapid screening of diverse bacteriophage libraries against SARS-CoV-2 Spike yielded unique Bicycle binders across the entire protein. Exploiting Bicycles' inherent chemical combinability, we converted early micromolar hits into nanomolar viral inhibitors through simple multimerization. We also show how combining Bicycles against different epitopes into a single biparatopic agent allows Spike from diverse variants of concern (VoC) to be targeted (Alpha, Beta, Delta and Omicron). Finally, we demonstrate in both male hACE2-transgenic mice and Syrian golden hamsters that both multimerized and biparatopic Bicycles reduce viraemia and prevent host inflammation. These results introduce Bicycles as a potential antiviral modality to tackle new and rapidly evolving viruses.
- Bicycle Therapeutics, Portway Building, Granta Park, Cambridge, CB21 6GS, United Kingdom.
Organizational Affiliation: