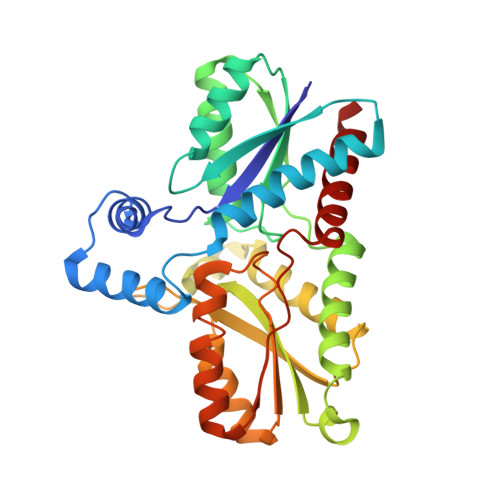
Active site architecture of coproporphyrin ferrochelatase with its physiological substrate coproporphyrin III: Propionate interactions and porphyrin core deformation.
Dali, A., Gabler, T., Sebastiani, F., Destinger, A., Furtmuller, P.G., Pfanzagl, V., Becucci, M., Smulevich, G., Hofbauer, S.(2023) Protein Sci 32: e4534-e4534
- PubMed: 36479958 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4534
- Primary Citation Related Structures:
8AT8, 8AW7 - PubMed Abstract:
Coproporphyrin ferrochelatases (CpfCs) are enzymes catalyzing the penultimate step in the coproporphyrin-dependent (CPD) heme biosynthesis pathway, which is mainly utilized by monoderm bacteria. Ferrochelatases insert ferrous iron into a porphyrin macrocycle and have been studied for many decades, nevertheless many mechanistic questions remain unanswered to date. Especially CpfCs, which are found in the CPD pathway, are currently in the spotlight of research. This pathway was identified in 2015 and revealed that the correct substrate for these ferrochelatases is coproporphyrin III (cpIII) instead of protoporphyrin IX, as believed prior the discovery of the CPD pathway. The chemistry of cpIII, which has four propionates, differs significantly from protoporphyrin IX, which features two propionate and two vinyl groups. These findings let us to thoroughly describe the physiological cpIII-ferrochelatase complex in solution and in the crystal phase. Here, we present the first crystallographic structure of the CpfC from the representative monoderm pathogen Listeria monocytogenes bound to its physiological substrate, cpIII, together with the in-solution data obtained by resonance Raman and UV-vis spectroscopy, for wild-type ferrochelatase and variants, analyzing propionate interactions. The results allow us to evaluate the porphyrin distortion and provide an in-depth characterization of the catalytically-relevant binding mode of cpIII prior to iron insertion. Our findings are discussed in the light of the observed structural restraints and necessities for this porphyrin-enzyme complex to catalyze the iron insertion process. Knowledge about this initial situation is essential for understanding the preconditions for iron insertion in CpfCs and builds the basis for future studies.
- Dipartimento di Chimica "Ugo Schiff" - DICUS, Università di Firenze, Sesto Fiorentino (FI), Italy.
Organizational Affiliation: