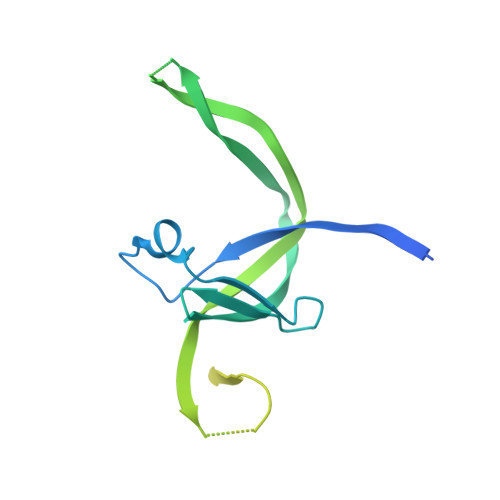
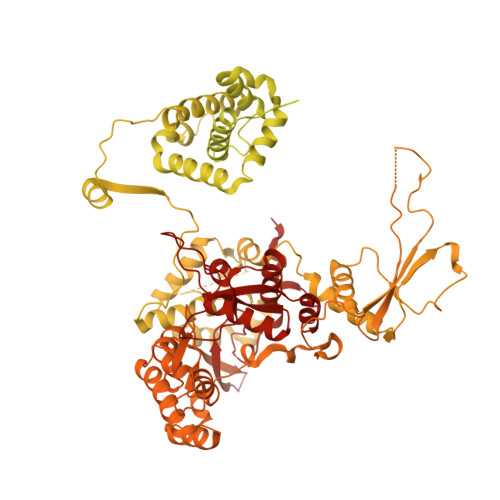
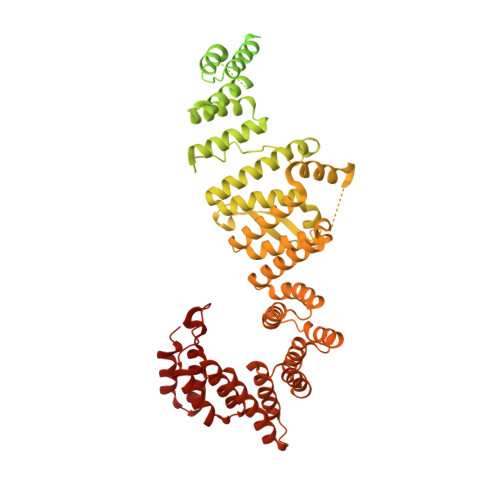
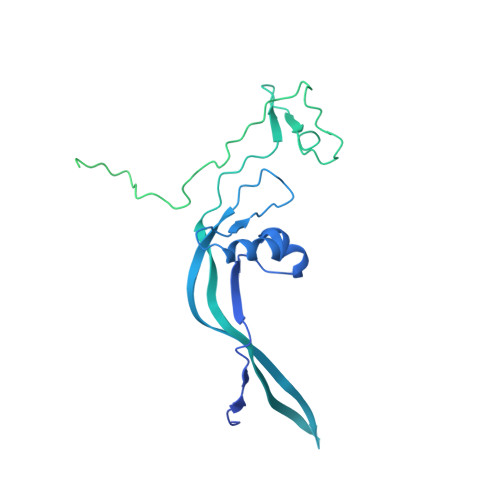
Structural insights into human TFIIIC promoter recognition.
Seifert-Davila, W., Girbig, M., Hauptmann, L., Hoffmann, T., Eustermann, S., Muller, C.W.(2023) Sci Adv 9: eadh2019-eadh2019
- PubMed: 37418517 Search on PubMed
- DOI: https://doi.org/10.1126/sciadv.adh2019
- Primary Citation Related Structures:
8CLI, 8CLJ, 8CLK, 8CLL - PubMed Abstract:
Transcription factor (TF) IIIC recruits RNA polymerase (Pol) III to most of its target genes. Recognition of intragenic A- and B-box motifs in transfer RNA (tRNA) genes by TFIIIC modules τA and τB is the first critical step for tRNA synthesis but is mechanistically poorly understood. Here, we report cryo-electron microscopy structures of the six-subunit human TFIIIC complex unbound and bound to a tRNA gene. The τB module recognizes the B-box via DNA shape and sequence readout through the assembly of multiple winged-helix domains. TFIIIC220 forms an integral part of both τA and τB connecting the two subcomplexes via a ~550-amino acid residue flexible linker. Our data provide a structural mechanism by which high-affinity B-box recognition anchors TFIIIC to promoter DNA and permits scanning for low-affinity A-boxes and TFIIIB for Pol III activation.
- Structural and Computational Biology Unit, European Molecular Biology Laboratory (EMBL), Meyerhofstrasse 1, 69117 Heidelberg, Germany.
Organizational Affiliation: