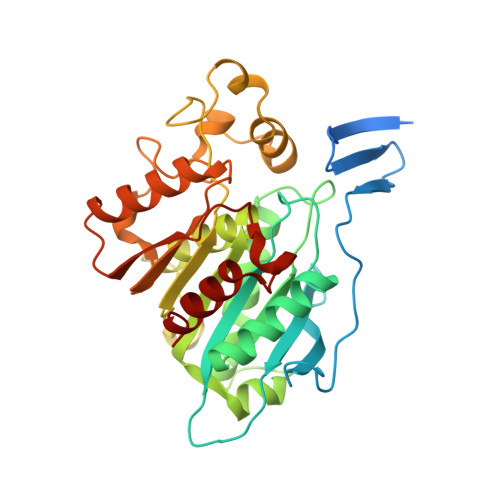
The protein conformational basis of isoflavone biosynthesis.
Wang, X., Pan, H., Sagurthi, S., Paris, V., Zhuo, C., Dixon, R.A.(2022) Commun Biol 5: 1249-1249
- PubMed: 36376429 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-022-04222-x
- Primary Citation Related Structures:
8E83, 8EA1, 8EA2 - PubMed Abstract:
Isoflavonoids play important roles in plant defense and also exhibit a range of mammalian health-promoting activities. Their biosynthesis is initiated by two enzymes with unusual catalytic activities; 2-hydroxyisoflavanone synthase (2-HIS), a membrane-bound cytochrome P450 catalyzing a coupled aryl-ring migration and hydroxylation, and 2-hydroxyisoflavanone dehydratase (2-HID), a member of a large carboxylesterase family that paradoxically catalyzes dehydration of 2-hydroxyisoflavanones to isoflavone. Here we report the crystal structures of 2-HIS from Medicago truncatula and 2-HID from Pueraria lobata. The 2-HIS structure reveals a unique cytochrome P450 conformation and heme and substrate binding mode that facilitate the coupled aryl-ring migration and hydroxylation reactions. The 2-HID structure reveals the active site architecture and putative catalytic residues for the dual dehydratase and carboxylesterase activities. Mutagenesis studies revealed key residues involved in substrate binding and specificity. Understanding the structural basis of isoflavone biosynthesis will facilitate the engineering of new bioactive isoflavonoids.
- BioDiscovery Institute and Department of Biological Sciences, University of North Texas, Denton, TX, 76203-5017, USA. Xiaoqiang.Wang@unt.edu.
Organizational Affiliation: