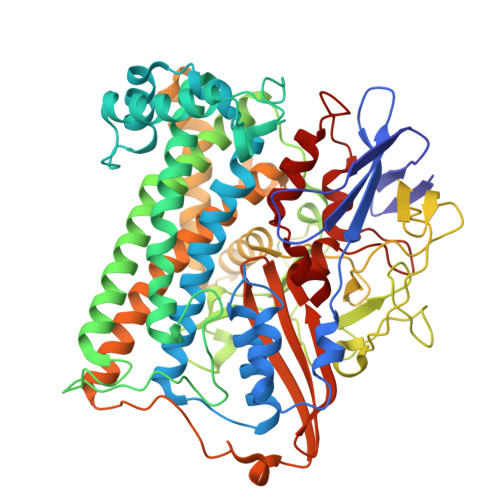
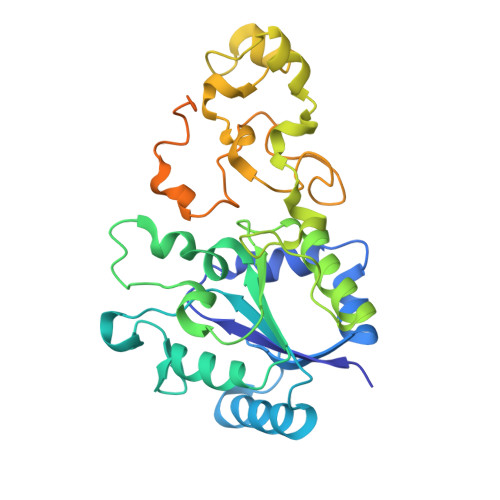
Stepwise conversion of the Cys 6 [4Fe-3S] to a Cys 4 [4Fe-4S] cluster and its impact on the oxygen tolerance of [NiFe]-hydrogenase.
Schmidt, A., Kalms, J., Lorent, C., Katz, S., Frielingsdorf, S., Evans, R.M., Fritsch, J., Siebert, E., Teutloff, C., Armstrong, F.A., Zebger, I., Lenz, O., Scheerer, P.(2023) Chem Sci 14: 11105-11120
- PubMed: 37860641 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d3sc03739h
- Primary Citation Related Structures:
8POU, 8POV, 8POW, 8POX, 8POY, 8POZ - PubMed Abstract:
The membrane-bound [NiFe]-hydrogenase of Cupriavidus necator is a rare example of a truly O 2 -tolerant hydrogenase. It catalyzes the oxidation of H 2 into 2e - and 2H + in the presence of high O 2 concentrations. This characteristic trait is intimately linked to the unique Cys 6 [4Fe-3S] cluster located in the proximal position to the catalytic center and coordinated by six cysteine residues. Two of these cysteines play an essential role in redox-dependent cluster plasticity, which bestows the cofactor with the capacity to mediate two redox transitions at physiological potentials. Here, we investigated the individual roles of the two additional cysteines by replacing them individually as well as simultaneously with glycine. The crystal structures of the corresponding MBH variants revealed the presence of Cys 5 [4Fe-4S] or Cys 4 [4Fe-4S] clusters of different architecture. The protein X-ray crystallography results were correlated with accompanying biochemical, spectroscopic and electrochemical data. The exchanges resulted in a diminished O 2 tolerance of all MBH variants, which was attributed to the fact that the modified proximal clusters mediated only one redox transition. The previously proposed O 2 protection mechanism that detoxifies O 2 to H 2 O using four protons and four electrons supplied by the cofactor infrastructure, is extended by our results, which suggest efficient shutdown of enzyme function by formation of a hydroxy ligand in the active site that protects the enzyme from O 2 binding under electron-deficient conditions.
- Charité - Universitätsmedizin Berlin, Corporate Member of Freie Universität Berlin and Humboldt-Universität zu Berlin, Institute of Medical Physics and Biophysics (CC2), Group Structural Biology of Cellular Signaling Charitéplatz 1 10117 Berlin Germany patrick.scheerer@charite.de.
Organizational Affiliation: