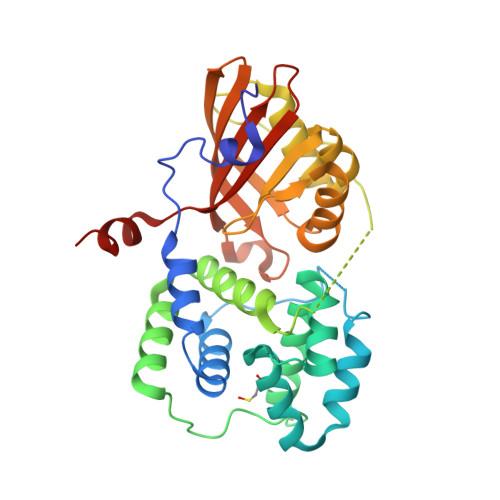
Structural framework for the understanding spectroscopic and functional signatures of the cyanobacterial Orange Carotenoid Protein families.
Sluchanko, N.N., Maksimov, E.G., Slonimskiy, Y.B., Varfolomeeva, L.A., Bukhanko, A.Y., Egorkin, N.A., Tsoraev, G.V., Khrenova, M.G., Ge, B., Qin, S., Boyko, K.M., Popov, V.O.(2024) Int J Biol Macromol 254: 127874-127874
- PubMed: 37939760 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2023.127874
- Primary Citation Related Structures:
8PYH, 8PZK - PubMed Abstract:
The Orange Carotenoid Protein (OCP) is a unique photoreceptor crucial for cyanobacterial photoprotection. Best studied Synechocystis sp. PCC 6803 OCP belongs to the large OCP1 family. Downregulated by the Fluorescence Recovery Protein (FRP) in low-light, high-light-activated OCP1 binds to the phycobilisomes and performs non-photochemical quenching. Recently discovered families OCP2 and OCP3 remain structurally and functionally underexplored, and no systematic comparative studies have ever been conducted. Here we present two first crystal structures of OCP2 from morphoecophysiologically different cyanobacteria and provide their comprehensive structural, spectroscopic and functional comparison with OCP1, the recently described OCP3 and all-OCP ancestor. Structures enable correlation of spectroscopic signatures with the effective number of hydrogen and discovered here chalcogen bonds anchoring the ketocarotenoid in OCP, as well as with the rotation of the echinenone's β-ionone ring in the CTD. Structural data also helped rationalize the observed differences in OCP/FRP and OCP/phycobilisome functional interactions. These data are expected to foster OCP research and applications in optogenetics, targeted carotenoid delivery and cyanobacterial biomass engineering.
- A.N. Bach Institute of Biochemistry, Federal Research Centre of Biotechnology of the Russian Academy of Sciences, Moscow 119071, Russia. Electronic address: nikolai.sluchanko@mail.ru.
Organizational Affiliation: