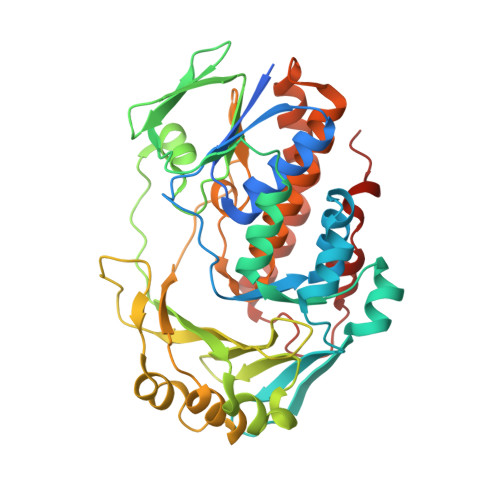
Ligand bound structure of a 6-hydroxynicotinic acid 3-monooxygenase provides mechanistic insights.
Turlington, Z.R., Vaz Ferreira de Macedo, S., Perry, K., Belsky, S.L., Faust, J.A., Snider, M.J., Hicks, K.A.(2024) Arch Biochem Biophys 752: 109859-109859
- PubMed: 38104959 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2023.109859
- Primary Citation Related Structures:
8UIQ, 8UIV - PubMed Abstract:
6-Hydroxynicotinic acid 3-monooxygenase (NicC) is a bacterial enzyme involved in the degradation of nicotinic acid. This enzyme is a Class A flavin-dependent monooxygenase that catalyzes a unique decarboxylative hydroxylation. The unliganded structure of this enzyme has previously been reported and studied using steady- and transient-state kinetics to support a comprehensive kinetic mechanism. Here we report the crystal structure of the H47Q NicC variant in both a ligand-bound (solved to 2.17 Å resolution) and unliganded (1.51 Å resolution) form. Interestingly, in the liganded form, H47Q NicC is bound to 2-mercaptopyridine (2-MP), a contaminant present in the commercial stock of 6-mercaptopyridine-3-carboxylic acid(6-MNA), a substrate analogue. 2-MP binds weakly to H47Q NicC and is not a substrate for the enzyme. Based on kinetic and thermodynamic characterization, we have fortuitously captured a catalytically inactive H47Q NicC•2-MP complex in our crystal structure. This complex reveals interesting mechanistic details about the reaction catalyzed by 6-hydroxynicotinic acid 3-monooxygenase.
- Department of Chemistry, State University of New York at Cortland, Cortland, NY, 13045, United States.
Organizational Affiliation: