Exploration of P1 and P4 modifications of nirmatrelvir: Design, synthesis, biological evaluation, and X-ray structural studies of SARS-CoV-2 Mpro inhibitors.
Ghosh, A.K., Yadav, M., Iddum, S., Ghazi, S., Lendy, E.K., Jayashankar, U., Beechboard, S.N., Takamatsu, Y., Hattori, S.I., Amano, M., Higashi-Kuwata, N., Mitsuya, H., Mesecar, A.D.(2024) Eur J Med Chem 267: 116132-116132
- PubMed: 38335815 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ejmech.2024.116132
- Primary Citation Related Structures:
8UND - PubMed Abstract:
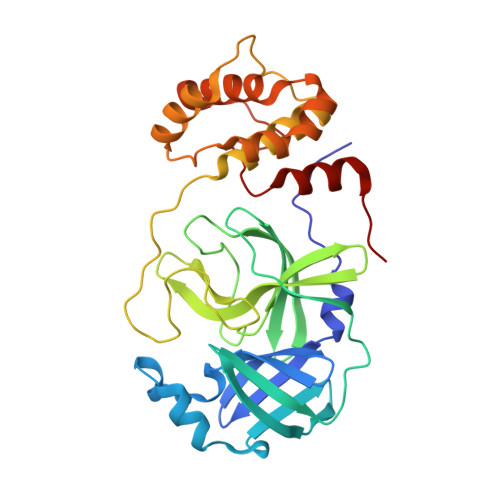
We report the synthesis, biological evaluation, and X-ray structural studies of a series of SARS-CoV-2 Mpro inhibitors based upon the X-ray crystal structure of nirmatrelvir, an FDA approved drug that targets the main protease of SARS-CoV-2. The studies involved examination of various P4 moieties, P1 five- and six-membered lactam rings to improve potency. In particular, the six-membered P1 lactam ring analogs exhibited high SARS-CoV-2 Mpro inhibitory activity. Several compounds effectively blocked SARS-CoV-2 replication in VeroE6 cells. One of these compounds maintained good antiviral activity against variants of concern including Delta and Omicron variants. A high-resolution X-ray crystal structure of an inhibitor bound to SARS-CoV-2 Mpro was determined to gain insight into the ligand-binding properties in the Mpro active site.
- Department of Chemistry, Purdue University, 560 Oval Drive, West Lafayette, IN, 47907, USA; Department of Medicinal Chemistry and Molecular Pharmacology, Purdue University, West Lafayette, IN, 47907, USA. Electronic address: akghosh@purdue.edu.
Organizational Affiliation: