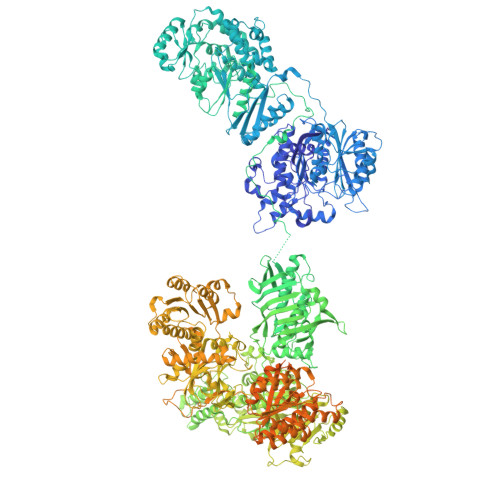
Snapshots of acyl carrier protein shuttling in human fatty acid synthase.
Schultz, K., Costa-Pinheiro, P., Gardner, L., Pinheiro, L.V., Ramirez-Solis, J., Gardner, S.M., Wellen, K.E., Marmorstein, R.(2025) Nature 641: 520-528
- PubMed: 39979457 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-025-08587-x
- Primary Citation Related Structures:
8VF7, 8VG4, 8VLE, 8VLO, 8VLP, 8VM0, 8VM5, 8VM6, 8VM7, 8VMC, 8VMD - PubMed Abstract:
The mammalian fatty acid synthase (FASN) enzyme is a dynamic multienzyme that belongs to the megasynthase family. In mammals, a single gene encodes six catalytically active domains and a flexibly tethered acyl carrier protein (ACP) domain that shuttles intermediates between active sites for fatty acid biosynthesis 1 . FASN is an essential enzyme in mammalian development through the role that fatty acids have in membrane formation, energy storage, cell signalling and protein modifications. Thus, FASN is a promising target for treatment of a large variety of diseases including cancer, metabolic dysfunction-associated fatty liver disease, and viral and parasite infections 2,3 . The multi-faceted mechanism of FASN and the dynamic nature of the protein, in particular of the ACP, have made it challenging to understand at the molecular level. Here we report cryo-electron microscopy structures of human FASN in a multitude of conformational states with NADPH and NADP + plus acetoacetyl-CoA present, including structures with the ACP stalled at the dehydratase (DH) and enoyl-reductase (ER) domains. We show that FASN activity in vitro and de novo lipogenesis in cells is inhibited by mutations at the ACP-DH and ACP-ER interfaces. Together, these studies provide new molecular insights into the dynamic nature of FASN and the ACP shuttling mechanism, with implications for developing improved FASN-targeted therapeutics.
- Abramson Family Cancer Research Institute, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA, USA. kollin.schultz@pennmedicine.upenn.edu.
Organizational Affiliation: