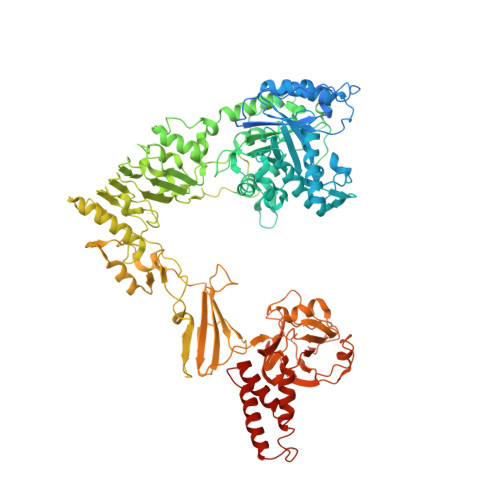
Structure-Based High-Efficiency Homogeneous Antibody Platform by Endoglycosidase Sz Provides Insights into Its Transglycosylation Mechanism.
Hsieh, Y.C., Guan, H.H., Lin, C.C., Huang, T.Y., Chuankhayan, P., Chen, N.C., Wang, N.H., Hu, P.L., Tsai, Y.C., Huang, Y.C., Yoshimura, M., Lin, P.J., Hsieh, Y.H., Chen, C.J.(2024) JACS Au 4: 2130-2150
- PubMed: 38938812
- DOI: https://doi.org/10.1021/jacsau.4c00004
- Primary Citation of Related Structures:
8W4G, 8W4I, 8W4L, 8W4M, 8W4N, 8X8G - PubMed Abstract:
Monoclonal antibodies (mAbs) have gradually dominated the drug markets for various diseases. Improvement of the therapeutic activities of mAbs has become a critical issue in the pharmaceutical industry. A novel endo-β- N -acetylglucosaminidase, EndoSz, from Streptococcus equi subsp. zooepidemicus Sz105 is discovered and applied to enhance the activities of mAbs. Our studies demonstrate that the mutant EndoSz-D234M possesses an excellent transglycosylation activity to generate diverse glycoconjugates on mAbs. We prove that EndoSz-D234M can be applied to various marketed therapeutic antibodies and those in development for antibody remodeling. The remodeled homogeneous antibodies (mAb-G2S2) produced by EndoSz-D234M increase the relative ADCC activities by 3-26-fold. We further report the high-resolution crystal structures of EndoSz-D234M in the apo -form at 2.15 Å and the complex form with a bound G2S2-oxazoline intermediate at 2.25 Å. A novel pH-jump method was utilized to obtain the complex structure with a high resolution. The detailed interactions of EndoSz-D234M and the carried G2S2-oxazoline are hence delineated. The oxazoline sits in a hole, named the oxa-hole, which stabilizes the G2S2-oxazoline in transit and catalyzes the further transglycosylation reaction while targeting Asn-GlcNAc (+1) of Fc. In the oxa-hole, the H-bonding network involved with oxazoline dominates the transglycosylation activity. A mobile loop2 (a.a. 152-159) of EndoSz-D234M reshapes the binding grooves for the accommodation of G2S2-oxazoline upon binding, at which Trp154 forms a hydrogen bond with Man (-2). The long loop4 (a.a. 236-248) followed by helix3 is capable of dominating the substrate selectivity of EndoSz-D234M. In addition, the stepwise transglycosylation behavior of EndoSz-D234M is elucidated. Based on the high-resolution structures of the apo -form and the bound form with G2S2-oxazoline as well as a systematic mutagenesis study of the relative transglycosylation activity, the transglycosylation mechanism of EndoSz-D234M is revealed.
- OBI Pharma, Inc., No. 508, Sec. 7, ZhongXiao E. Rd, Nangang Dist., Taipei City 115, Taiwan.
Organizational Affiliation: