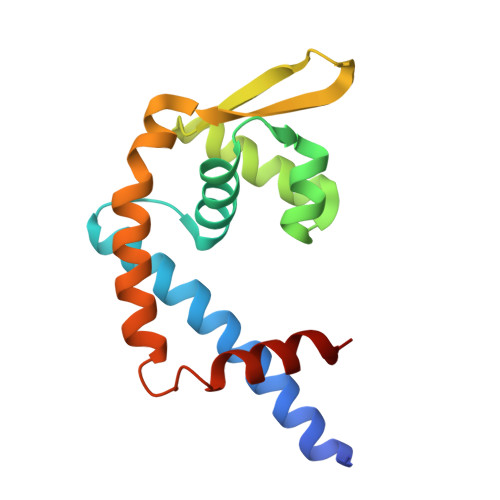
Crystal structure of a Clostridioides difficile multiple antibiotic resistance regulator (MarR) CD0473 suggests a potential redox-regulated function.
Kwon, N., Rho, S., Ha, S.C., Park, S.(2024) Int J Biol Macromol 280: 136036-136036
- PubMed: 39332572 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2024.136036
- Primary Citation Related Structures:
8XT8, 8XTA, 8XU0 - PubMed Abstract:
Clostridioides difficile may constitute a small part of normal gut microbiota in humans without causing any symptoms, but an uncontrolled growth common to hospitalized patients can cause Clostridioides difficile infection (CDI) leading to severe colonic symptoms. As the bacteria are attaining resistance to various antibiotics worldwide, CDI is becoming a serious public health problem. Although a family of transcription factors called MarR (Multiple antibiotic resistance Regulator) plays a key role in the bacterial response to various environmental stresses including antibiotics, most of the 14 MarRs predicted to exist in the C. difficile genome lack structural or functional studies. In this respect, X-ray crystal structure of a C. difficile MarR CD0473 with a yet unknown function has been determined using a Hg-soaked crystal. In the structure, two closely located flexible conformations of Hg-bound cysteines suggested a possibility of intra-subunit disulfide bridge formation. By searching the neighboring intergenic regions of CD0473, two pseudo-palindromic DNA sites were found and shown to bind the protein. MarR CD0473 binding stronger to the DNA in an oxidizing condition supported further that it may function as a redox regulated switch likely via its oxidized disulfide formation.
- School of Systems Biomedical Science and Integrative Institute of Basic Sciences, Soongsil University, Seoul 06978, Republic of Korea.
Organizational Affiliation: