A photo-SAR study of photoswitchable azobenzene tubulin-inhibiting antimitotics identifying a general method for near-quantitative photocontrol.
Reynders, M., Garscia, M., Muller-Deku, A., Wranik, M., Krauskopf, K., de la Osa de la Rosa, L., Schaffer, K., Jotten, A., Rode, A., Stierle, V., Kraus, Y., Baumgartner, B., Ali, A., Bubeneck, A., Seal, T., Steinmetz, M.O., Paulitschke, P., Thorn-Seshold, O.(2024) Chem Sci 15: 12301-12309
- PubMed: 39118608 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d4sc03072a
- Primary Citation Related Structures:
9F8G - PubMed Abstract:
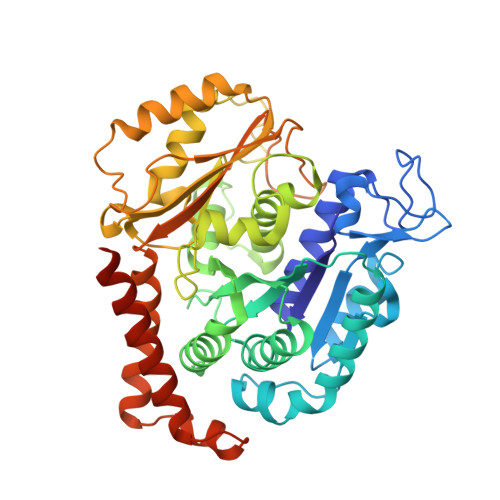
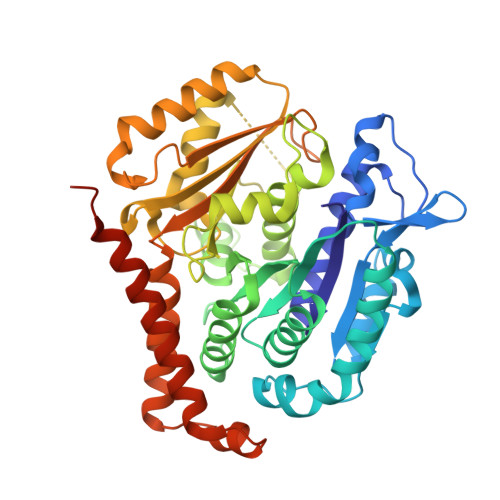
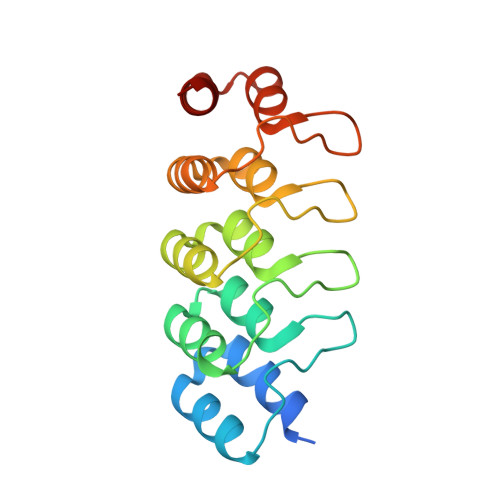
Azobenzene analogues of the tubulin polymerisation inhibitor combretastatin A4 (PSTs) were previously developed to optically control microtubule dynamics in living systems, with subsecond response time and single-cell spatial precision, by reversible in situ photoswitching of their bioactivity with near-UV/visible light. First-generation PSTs were sufficiently potent and photoswitchable for use in live cells and embryos. However, the link between their seconds-scale and hours-scale bioactivity remained untested. Furthermore, the scope for modifications to tune their photo-structure-activity-relationship or expand their function was unknown. Here, we used large-field-of-view, long-term tandem photoswitching/microscopy to reveal the temporal onset of cytostatic effects. We then synthesised a panel of novel PSTs exploring structural variations that tune photoresponse wavelengths and lipophilicity, identifying promising blue-shifted analogues that are better-compatible with GFP/YFP imaging. Taken together, these results can guide new design and applications for photoswitchable microtubule inhibitors. We also identified tolerated sites for linkers to attach functional cargos; and we tested fluorophores, aiming at RET isomerisation or reporter probes. Instead we found that these antennas greatly enhance long-wavelength single-photon photoisomerisation, by an as-yet un-explored mechanism, that can now drive general progress towards near-quantitative long-wavelength photoswitching of photopharmaceuticals in living systems, with minimal molecular redesign and broad scope.
- Faculty of Chemistry and Pharmacy, Ludwig-Maximilians-University Munich Munich 81377 Germany oliver.thorn-seshold@cup.lmu.de.
Organizational Affiliation: