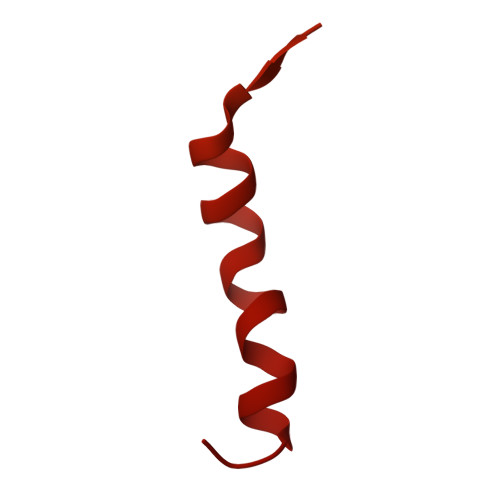
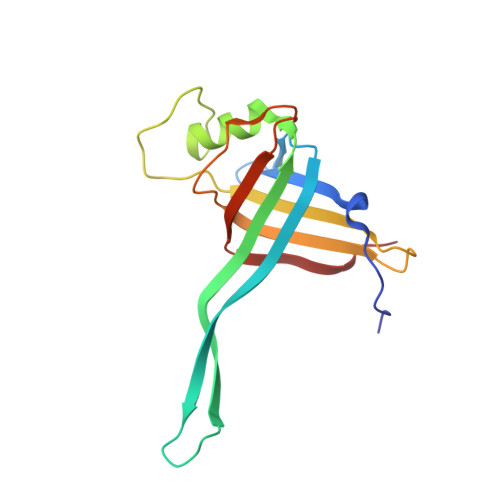
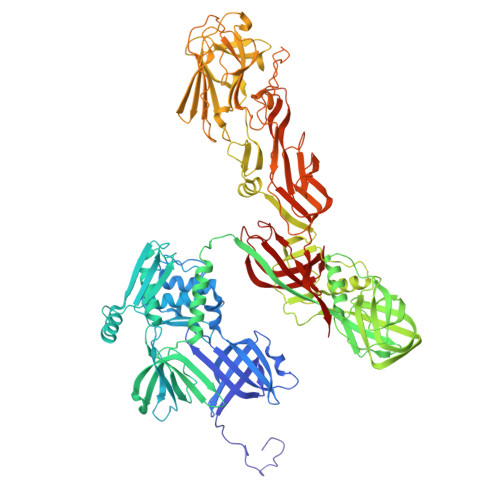
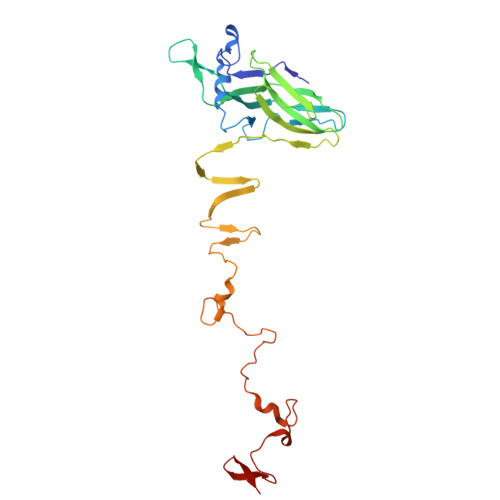
Cryo-EM resolves the structure of the archaeal dsDNA virus HFTV1 from head to tail.
Zhang, D.X., Isupov, M.N., Davies, R.M., Schwarzer, S., McLaren, M., Stuart, W.S., Gold, V.A.M., Oksanen, H.M., Quax, T.E.F., Daum, B.(2025) Sci Adv 11: eadx1178-eadx1178
- PubMed: 41042861 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adx1178
- Primary Citation Related Structures:
8QPG, 8QPQ, 8QQN, 8QSI, 8QSY, 9FKB, 9GS0, 9H4P, 9H5B, 9H7V - PubMed Abstract:
While archaeal viruses show a stunning diversity of morphologies, many bear a notable resemblance to tailed bacterial phages. This raises fundamental questions: Do all tailed viruses share a common origin and do they infect their hosts in similar ways? Answering these questions requires high-resolution structural insights, yet no complete atomic models of archaeal viruses have been available. Here, we present the near-atomic resolution structure of Haloferax tailed virus 1 (HFTV1), an archaeal virus thriving in extreme salinity. Using cryo-electron microscopy, we resolve the architecture and assembly of all structural proteins and capture conformational transitions associated with DNA ejection. Our data reveal genome spooling within the capsid and identify putative receptor-binding and catalytic sites for host recognition and infection. These findings uncover key mechanisms of archaeal virus assembly, principles of virus-host interactions, and evolutionary links connecting archaeal, bacterial, and eukaryotic viruses.
- Living Systems Institute, University of Exeter, Exeter EX4 4QD, UK.
Organizational Affiliation: