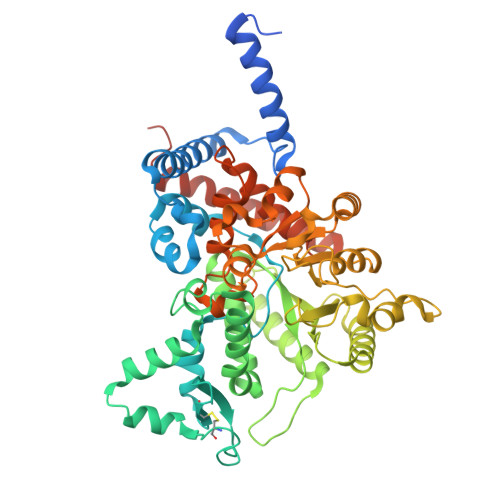
Structural basis for the substrate specificity of Helix pomatia AMP deaminase and a chimeric ADGF adenosine deaminase.
Kaur, G., Horton, J.R., Tzertzinis, G., Zhou, J., Schildkraut, I., Cheng, X.(2025) J Biological Chem 301: 110357-110357
- PubMed: 40505866 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2025.110357
- Primary Citation Related Structures:
9NTE, 9NTF, 9NTG, 9NTH, 9NTI, 9NTJ, 9NTK - PubMed Abstract:
Helix pomatia AMP deaminase (HPAMPD), an enzyme enriched in the foot muscle of the mollusk Helix pomatia, exhibits deaminase activity on adenosine-5'-monophosphate (AMP). HPAMPD is the first member of the adenosine deaminase-related growth factor (ADGF) family to prefer the nucleotideAMP over the nucleoside adenosine. To investigate the substrate selectivity of HPAMPD, we determined its structure in both the apo form and in complex with the adenosine analogs pentostatin and pentostatin-5'-monophosphate. Structurally, HPAMPD adopts a fold similar to human ADA2, an ADGF family member. HPAMPD has acquired the ability to interact with the 5'-monophosphate group of AMP through polar and charged residues located in three key structural elements: (1) the loop immediately following strand β1; (2) the loop between helices αH and αI; and (3) the end of strand β5 and its adjacent loop. We engineered a chimeric deaminase by integrating these elements from HPAMPD into another related mollusk nucleoside adenosine deaminase, Aplysia ADGF. The chimeric enzyme efficiently deaminates AMP, demonstrating a gained substrate specificity, while retaining the adenosine deamination activity of Aplysia ADGF. The phosphate-binding feature of HPAMPD is a hallmark of nucleotide deaminases, conserved among AMP and N6-methyl-AMP (6mAMP) deaminases. We discuss the human adenosine deaminases each with distinct substrate specificities for the nucleoside, the nucleotide (AMP), and its methylated form, 6mAMP.
- Department of Epigenetics and Molecular Carcinogenesis, The University of Texas MD Anderson Cancer Center, Houston, TX 77030, USA.
Organizational Affiliation: