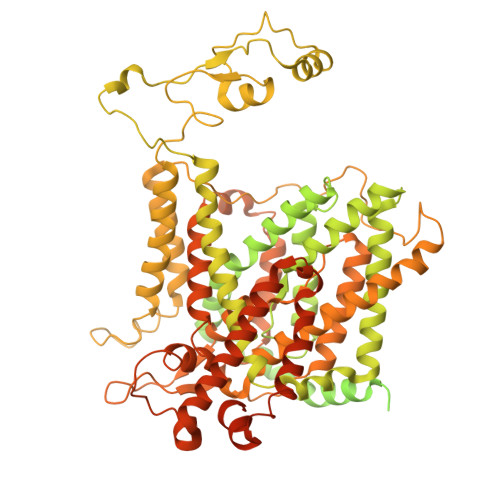
CryoEM and computational modeling structural insights into the pH regulator NBCn1.
Wang, W., R Zhekova, H., Tsirulnikov, K., Dwadasi, B.S., Aparicio, E.G., Azimov, R., Abuladze, N., Kao, L., Acuna, D., Tieleman, D.P., Zhou, Z.H., Pushkin, A., Kurtz, I.(2025) Nat Commun 16: 9932-9932
- PubMed: 41219199 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-64868-z
- Primary Citation Related Structures:
9OVR - PubMed Abstract:
Breast cancer cells survive despite being exposed to a toxic acidic extracellular environment, by utilizing the NBCn1 transporter. The molecular basis for this phenomenon is unknown, given the lack of an NBCn1 atomic structural model. We therefore determined the 3.3 Å cryoEM structure of the human NBCn1 outward facing (OF) conformational state with densities corresponding to the transported ions in the ion coordination site. We further generated NBCn1 inward facing (IF) and intermediate (occluded) structures and characterized the transport cycle and the ion dynamics in the IF and OF states. The results showed that NBCn1 utilizes an elevator-type transport mechanism with a small vertical shift of the ion coordination site between OF and IF conformational states and that the transported ions permeate without significant energy barriers. Functional experiments showed that NBCn1 has an extremely high ion turnover rate (TOR) of ~15,000 s -1 . The unusually high NBCn1 TOR value associated with the small protein structural changes during the OF to IF transitions and the favorable ion permeation energetics provides breast cancer cells with a highly efficient base loading mechanism contributing to their survival advantage.
- Department of Medicine, Division of Nephrology, David Geffen School of Medicine, University of California, Los Angeles, CA, USA.
Organizational Affiliation: