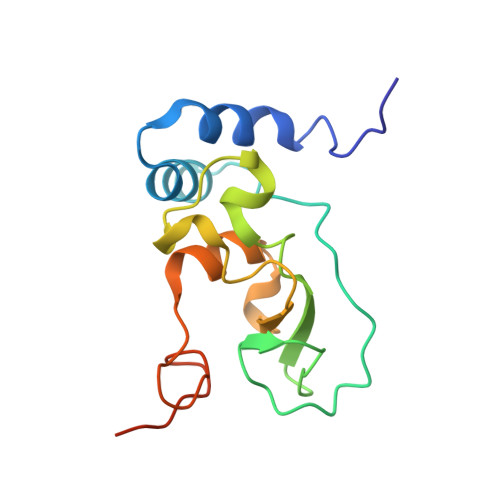
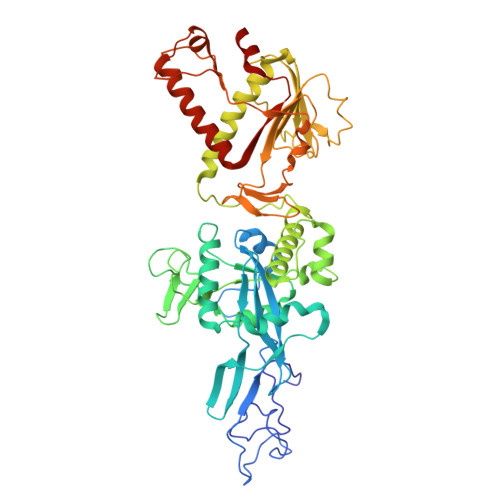
Structural and catalytic diversity of coronavirus proofreading exoribonuclease.
Li, Y., Cao, X., Recker, L.M., Yang, Y., Liu, C.(2025) Nat Commun 17: 452-452
- PubMed: 41361186
- DOI: https://doi.org/10.1038/s41467-025-67140-6
- Primary Citation of Related Structures:
9YCK, 9YCL, 9YCM, 9YCN, 9YCO - PubMed Abstract:
The coronavirus proofreading exoribonuclease (ExoN) is essential for genome fidelity and immune evasion of the viruses. Despite its critical roles in the viral life cycle, it is unclear how ExoNs across different coronaviruses diverge in their structures and catalytic properties, which may lead to differences in viral genome mutation rates and, consequently, viral fitness, immune evasion, and resistance to antiviral drugs. Here, we present comparative structural and biochemical analyses of ExoNs between two most representative human coronaviruses, Middle East Respiratory Syndrome Coronavirus (MERS-CoV) from the merbecovirus subgenus and SARS-CoV-2 from the sarbecovirus subgenus. Our results reveal a markedly lower catalytic activity of ExoN from MERS-CoV than that from SARS-CoV-2. The molecular basis of such a divergence across the two coronaviruses is unveiled by the cryo-EM structures of MERS-CoV ExoN in complex with RNA substrates bearing different 3'-end base pairs or mismatch, which represent the first set of ExoN structures from a coronavirus outside the sarbecovirus subgenus. Our findings also identify two highly conserved structural determinants that dictate efficient excision of different nucleotides at the 3' terminus of RNA substrates by coronavirus ExoNs, a property that is pivotal for their roles in both viral RNA proofreading and immune evasion.
- Roy J. Carver Department of Biochemistry, Biophysics and Molecular Biology, Iowa State University, Ames, IA, USA.
Organizational Affiliation: