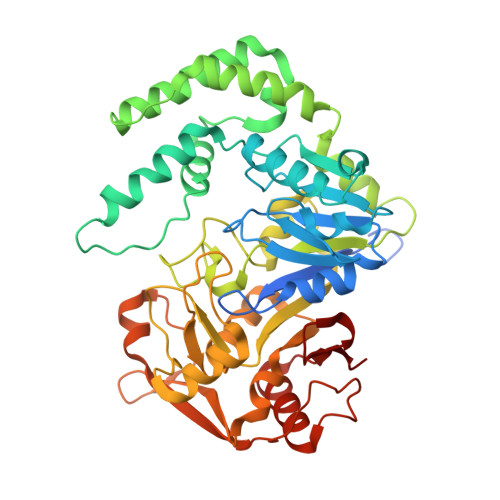
Recombinant mouse muscle adenylosuccinate synthetase: overexpression, kinetics, and crystal structure.
Iancu, C.V., Borza, T., Choe, J.Y., Fromm, H.J., Honzatko, R.B.(2001) J Biological Chem 276: 42146-42152
- PubMed: 11560929 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M106294200
- Primary Citation Related Structures:
1J4B - PubMed Abstract:
Vertebrates possess two isozymes of adenylosuccinate synthetase. The acidic isozyme is similar to the synthetase from bacteria and plants, being involved in the de novo biosynthesis of AMP, whereas the basic isozyme participates in the purine nucleotide cycle. Reported here is the first instance of overexpression and crystal structure determination of a basic isozyme of adenylosuccinate synthetase. The recombinant mouse muscle enzyme purified to homogeneity in milligram quantities exhibits a specific activity comparable with that of the rat muscle enzyme isolated from tissue and K(m) parameters for GTP, IMP, and l-aspartate (12, 45, and 140 microm, respectively) similar to those of the enzyme from Escherichia coli. The mouse muscle and E. coli enzymes have similar polypeptide folds, differing primarily in the conformation of loops, involved in substrate recognition and stabilization of the transition state. Residues 65-68 of the muscle isozyme adopt a conformation not observed in any previous synthetase structure. In its new conformation, segment 65-68 forms intramolecular hydrogen bonds with residues essential for the recognition of IMP and, in fact, sterically excludes IMP from the active site. Observed differences in ligand recognition among adenylosuccinate synthetases may be due in part to conformational variations in the IMP pocket of the ligand-free enzymes.
- Department of Biochemistry, Biophysics, and Molecular Biology, Molecular Biology Building, Iowa State University, Ames, Iowa 5011, USA.
Organizational Affiliation: