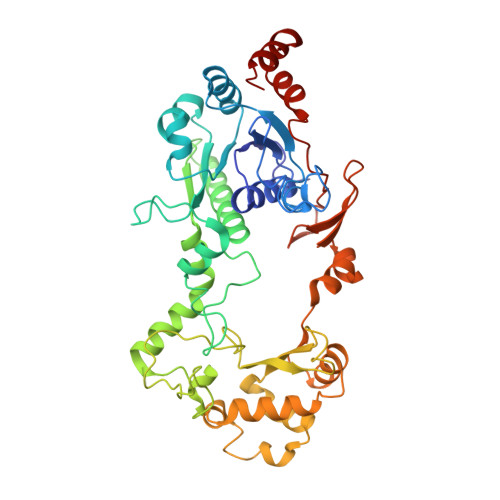
Formyl-CoA Transferase encloses the CoA binding site at the interface of an interlocked dimer
Ricagno, S., Jonsson, S., Richards, N., Lindqvist, Y.(2003) EMBO J 22: 3210-3219
- PubMed: 12839984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg333
- Primary Citation Related Structures:
1P5H, 1P5R - PubMed Abstract:
Formyl-CoA transferase catalyses transfer of CoA from formate to oxalate in the first step of oxalate degradation by Oxalobacter formigenes, a bacterium present in the intestinal flora which is implicated in oxalate catabolism in mammals. Formyl-CoA transferase is a member of a family of CoA-transferases for which no structural information is available. We now report the three-dimensional structure of O.formigenes formyl-CoA transferase, which reveals a novel fold and a very striking assembly of the homodimer. The subunit is composed of a large and a small domain where residues from both the N- and C-termini of the subunit are part of the large domain. The linkers between the domains give the subunit a circular shape with a hole in the middle. The enzyme monomers are tightly interacting and are interlocked. This fold requires drastic rearrangement of approximately 75 residues at the C-terminus for formation of the dimer. The structure of a complex of formyl-CoA transferase with CoA is also reported and sets the scene for a mechanistic understanding of enzymes of this family of CoA-transferases.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-17177 Stockholm, Sweden.
Organizational Affiliation: