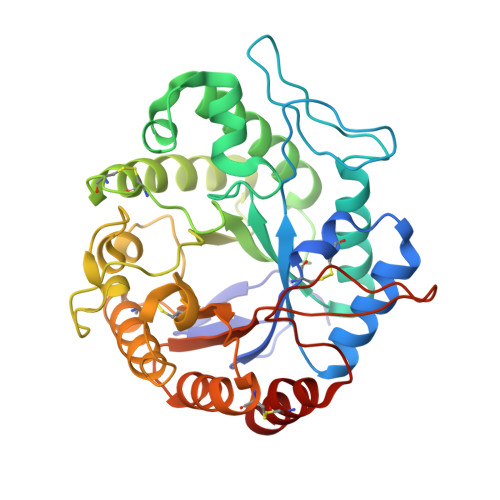
The Three-Dimensional Structure of a Trichoderma Reesei Beta-Mannanase from Glycoside Hydrolase Family 5.
Sabini, E., Schubert, H., Murshudov, G., Wilson, K.S., Siika-Aho, M., Penttila, M.(2000) Acta Crystallogr D Biol Crystallogr 56: 3
- PubMed: 10666621 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444999013943
- Primary Citation Related Structures:
1QNO, 1QNP, 1QNQ, 1QNR, 1QNS - PubMed Abstract:
The crystal structure of the catalytic core domain of beta-mannanase from the fungus Trichoderma reesei has been determined at a resolution of 1.5 A. The structure was solved using the anomalous scattering from a single non-isomorphous platinum complex with two heavy-metal sites in space group P2(1). The map computed with the experimental phases was enhanced by the application of an automated model building and refinement procedure using the amplitudes and experimental phases as observations. This approach is expected to be of more general application. The structure of the native enzyme and complexes with Tris-HCl and mannobiose are also reported: the mannobiose binds in subsites +1 and +2. The structure is briefly compared with that of the homologous beta-mannanase from the bacterium Thermomonospora fusca.
- Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5DD, England.
Organizational Affiliation: