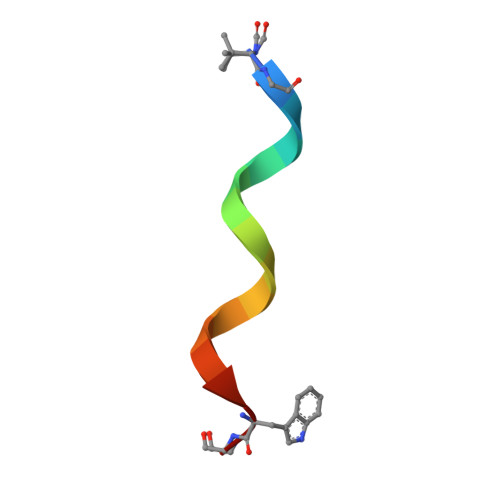
Membrane Protein Crystallization in Lipidic Mesophases. Hosting Lipid Effects on the Crystallization and Structure of a Transmembrane Peptide
Hoefer, N., Aragao, D., Lyons, J., Caffrey, M.(2011) Cryst Growth Des 11: 1182
- PubMed: 22933857 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/cg101384p
- Primary Citation Related Structures:
2Y5M, 2Y6N - PubMed Abstract:
Gramicidin is an apolar pentadecapeptide antibiotic consisting of alternating D-and L-amino acids. It functions, in part, by creating pores in membranes of susceptible cells rendering them leaky to monovalent cations. The peptide should be able to traverse the host membrane either as a double stranded, intertwined double helix (DSDH) or as a head-to-head single stranded helix (HHSH). Current structure models are based on macromolecular X-ray crystallography (MX) and nuclear magnetic resonance (NMR). However, the HHSH form has only been observed by NMR. The shape and size of the different gramicidin conformations differ. We speculated therefore that reconstituting it into a lipidic mesophase with bilayers of different microstructures would preferentially stabilize one form over the other. By using such mesophases for in meso crystallogenesis the expectation was that at least one would generate crystals of gramicidin in the HHSH form for structure determination by MX. This was tested using commercial and in-house synthesised lipids that support in meso crystallogenesis. Lipid acyl chain lengths were varied from 14 to 18 carbons to provide mesophases with a range of bilayer thicknesses. Unexpectedly, all lipids produced high quality, structure-grade crystals with gramicidin only in the DSDH conformation.
- Membrane Structural and Functional Biology Group, School of Biochemistry and Immunology, and School of Medicine, Trinity College, Dublin.
Organizational Affiliation: