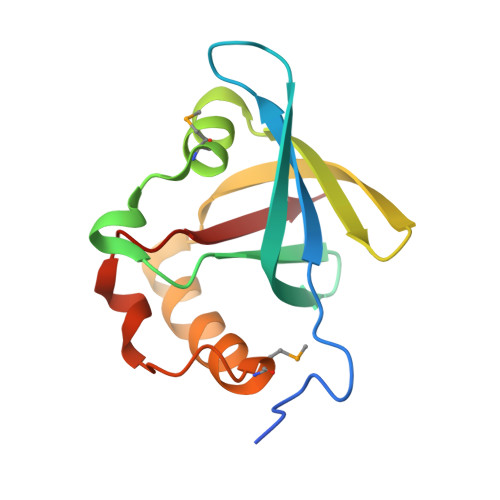
Structure of Ddn, the deazaflavin-dependent nitroreductase from Mycobacterium tuberculosis involved in bioreductive activation of PA-824.
Cellitti, S.E., Shaffer, J., Jones, D.H., Mukherjee, T., Gurumurthy, M., Bursulaya, B., Boshoff, H.I., Choi, I., Nayyar, A., Lee, Y.S., Cherian, J., Niyomrattanakit, P., Dick, T., Manjunatha, U.H., Barry, C.E., Spraggon, G., Geierstanger, B.H.(2012) Structure 20: 101-112
- PubMed: 22244759 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2011.11.001
- Primary Citation Related Structures:
3R5L, 3R5P, 3R5R, 3R5W, 3R5Y, 3R5Z - PubMed Abstract:
Tuberculosis continues to be a global health threat, making bicyclic nitroimidazoles an important new class of therapeutics. A deazaflavin-dependent nitroreductase (Ddn) from Mycobacterium tuberculosis catalyzes the reduction of nitroimidazoles such as PA-824, resulting in intracellular release of lethal reactive nitrogen species. The N-terminal 30 residues of Ddn are functionally important but are flexible or access multiple conformations, preventing structural characterization of the full-length, enzymatically active enzyme. Several structures were determined of a truncated, inactive Ddn protein core with and without bound F(420) deazaflavin coenzyme as well as of a catalytically competent homolog from Nocardia farcinica. Mutagenesis studies based on these structures identified residues important for binding of F(420) and PA-824. The proposed orientation of the tail of PA-824 toward the N terminus of Ddn is consistent with current structure-activity relationship data.
- Genomics Institute of the Novartis Research Foundation, 10675 John Jay Hopkins Drive, San Diego, CA 92121-1125, USA.
Organizational Affiliation: