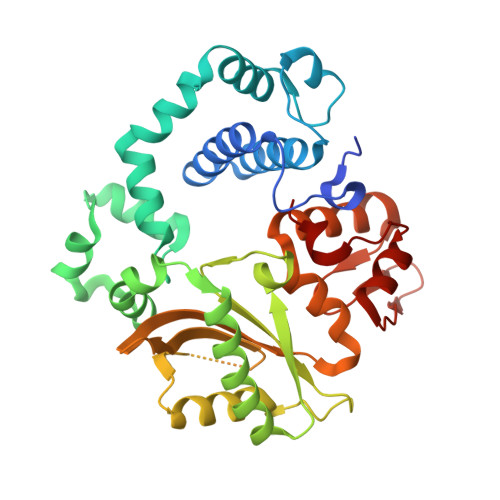
Watching a double strand break repair polymerase insert a pro-mutagenic oxidized nucleotide.
Jamsen, J.A., Sassa, A., Shock, D.D., Beard, W.A., Wilson, S.H.(2021) Nat Commun 12: 2059-2059
- PubMed: 33824325 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-21354-6
- Primary Citation Related Structures:
7KSS, 7KST, 7KSU, 7KSV, 7KSW, 7KSX, 7KSY, 7KSZ, 7KT0, 7KT1, 7KT2, 7KT3, 7KT4, 7KT5, 7KT6, 7KT7, 7KT8, 7KT9, 7KTA, 7KTB, 7KTC, 7KTD, 7KTE, 7KTF, 7KTG, 7KTH, 7KTI, 7KTJ, 7KTK, 7KTL, 7KTM, 7KTN - PubMed Abstract:
Oxidized dGTP (8-oxo-7,8-dihydro-2´-deoxyguanosine triphosphate, 8-oxodGTP) insertion by DNA polymerases strongly promotes cancer and human disease. How DNA polymerases discriminate against oxidized and undamaged nucleotides, especially in error-prone double strand break (DSB) repair, is poorly understood. High-resolution time-lapse X-ray crystallography snapshots of DSB repair polymerase μ undergoing DNA synthesis reveal that a third active site metal promotes insertion of oxidized and undamaged dGTP in the canonical anti-conformation opposite template cytosine. The product metal bridged O8 with product oxygens, and was not observed in the syn-conformation opposite template adenine (A t ). Rotation of A t into the syn-conformation enabled undamaged dGTP misinsertion. Exploiting metal and substrate dynamics in a rigid active site allows 8-oxodGTP to circumvent polymerase fidelity safeguards to promote pro-mutagenic double strand break repair.
- Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, NC, 27709, USA.
Organizational Affiliation: