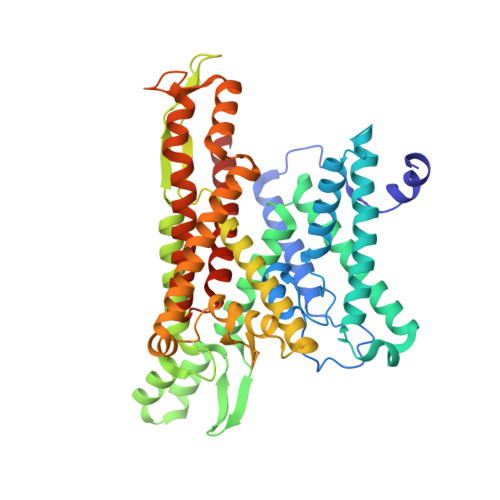
A common mechanism of Sec61 translocon inhibition by small molecules.
Itskanov, S., Wang, L., Junne, T., Sherriff, R., Xiao, L., Blanchard, N., Shi, W.Q., Forsyth, C., Hoepfner, D., Spiess, M., Park, E.(2023) Nat Chem Biol 19: 1063-1071
- PubMed: 37169959 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-023-01337-y
- Primary Citation Related Structures:
8DNV, 8DNW, 8DNX, 8DNY, 8DNZ, 8DO0, 8DO1, 8DO2, 8DO3 - PubMed Abstract:
The Sec61 complex forms a protein-conducting channel in the endoplasmic reticulum membrane that is required for secretion of soluble proteins and production of many membrane proteins. Several natural and synthetic small molecules specifically inhibit Sec61, generating cellular effects that are useful for therapeutic purposes, but their inhibitory mechanisms remain unclear. Here we present near-atomic-resolution structures of human Sec61 inhibited by a comprehensive panel of structurally distinct small molecules-cotransin, decatransin, apratoxin, ipomoeassin, mycolactone, cyclotriazadisulfonamide and eeyarestatin. All inhibitors bind to a common lipid-exposed pocket formed by the partially open lateral gate and plug domain of Sec61. Mutations conferring resistance to the inhibitors are clustered at this binding pocket. The structures indicate that Sec61 inhibitors stabilize the plug domain in a closed state, thereby preventing the protein-translocation pore from opening. Our study provides the atomic details of Sec61-inhibitor interactions and the structural framework for further pharmacological studies and drug design.
- Biophysics Graduate Program, University of California, Berkeley, Berkeley, CA, USA.
Organizational Affiliation: